LONDON — As the smallest living organisms on Earth, bacteria are difficult to study, even with today’s advancements in technology. Still, research on these microorganisms is crucial for the development of certain drugs, such as antibiotics. To better understand the antibiotic-resistant capabilities of some species, scientists from the U.S. and U.K. worked to develop the clearest image to date of the surface of a bacterial cell.
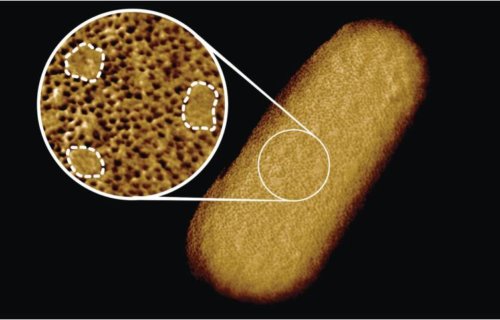
The study of the bacterium Escherichia coli (E. coli), reveals tightly compacted sections of proteins on the membrane, as well as sections that seem to lack proteins. Researchers say that the protein-lacking areas actually contained glycolipids, which function to keep the rigid structure of the membrane and signal other outside molecules.
As a gram-negative species, E. coli is known for its resistance to antibiotics. Like many other bacterial species, the outer membrane does not allow certain substances to infiltrate the cell, which causes dangerous diseases. “The outer membrane is a formidable barrier against antibiotics and is an important factor in making infectious bacteria resistant to medical treatment. However, it remains relatively unclear how this barrier is put together, which is why we chose to study it in such detail,” explains corresponding author Bart Hoogenboom, a professor of physics and astronomy at the University College London, in a statement.
“By studying live bacteria from the molecular to cellular scale, we can see how membrane proteins form a network that spans the entire surface of the bacteria, leaving small gaps for patches that contain no protein. This suggests that the barrier may not be equally hard to breach or stretch all over the bacterium, but may have stronger and weaker spots that can also be targeted by antibiotics.”
A microscopic needle “felt” the bacterium’s surface, outlining the molecules present. The outline showed dense areas of proteins that act as guards for the cell, only allowing beneficial substances in while restricting harmful substances. The gaps, filled with glycolipids, were sporadic throughout the protein patches, and only a few were detected. This suggests the essentially impenetrable surface of the bacteria stems from such a dense network of “guard” proteins.
“The textbook picture of the bacterial outer membrane shows proteins distributed over the membrane in a disordered manner, well-mixed with other building blocks of the membrane. Our images demonstrate that that is not the case, but that lipid patches are segregated from protein-rich networks just like oil separating from water, in some cases forming chinks in the armor of the bacteria,” notes Georgina Benn, who was responsible for the image of the bacterium taken in Hoogenboom’s lab at UCL.
E.coli’s rigid, dense membrane enables them to resist antibiotics, however, they are still capable of growth at an extremely fast pace. Researchers hypothesize that the sections of glycolipids provide the flexibility needed for the cells to grow and reproduce. “This new way of looking at the outer membrane means that we can now start exploring if and how such order matters for membrane function, integrity, and resistance to antibiotics,” concludes Benn.
This study is published in the Proceedings of the National Academy of Sciences.