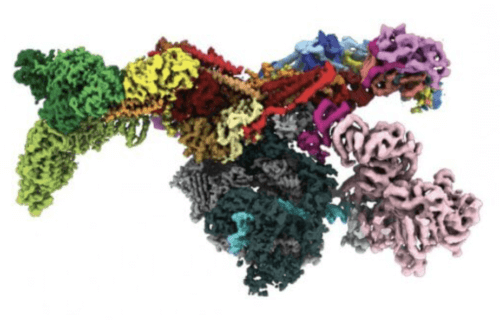
EVANSTON, Ill. — Scientists have created — for the first time ever — a 3D image of the structure inside human cells that are behind gene expression.
According to the National Human Genome Research Institute, gene expression “is the process by which the information encoded in a gene is used to direct the assembly of a protein molecule. The cell reads the sequence of the gene in groups of three bases. Each group of three bases (codon) corresponds to one of 20 different amino acids used to build the protein.”
Since the late 1800s geneticists have studied the phenomenon known as gene regulation, which essentially oversees gene expression. The process requires specialized proteins, however, the mechanism by which these proteins turn genes on and off has largely remained a mystery. In a recent study, however, a group of researchers from Northwestern University obtained a 3D image of the complex gene-regulator protein referred to as the Mediator-bound pre-initiation complex (Med-PIC).
Made up of 56 subunits, the large polypeptide nicknamed the “Mediator” helps to regulate gene expression in virtually every cell by placing the correct proteins in the appropriate locations for transcription, the first step to forming a specific protein.
“This machine is so basic to every branch of modern molecular biology in the context of gene expression,” explains senior author Yuan He, assistant professor of molecular biosciences in Northwestern’s Weinberg College of Arts and Sciences, in a statement. “Visualizing the structure in 3D will help us answer basic biological questions, such as how DNA is copied to RNA.”
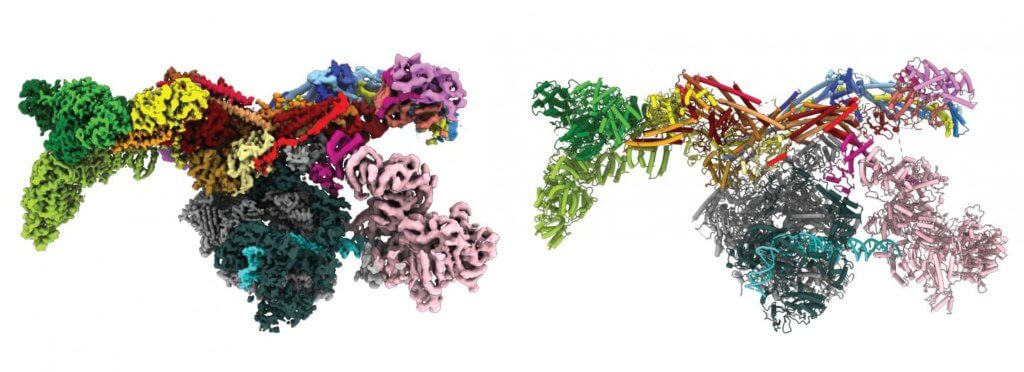
Though it was first identified in 1990, scientists were unable to visualize its full molecular structure, making it difficult to ascertain its physiology. Using cryogenic electron microscopy (cryo-EM), the team was able to gain a better insight into the method by which the protein functions. Since this protein is implicated in a wide variety of illnesses, such as cancer, neurological diseases, and HIV, the teams’ improved knowledge of its composition may one day be used to develop novel treatments.
“Seeing this structure allows us to understand how it works. It’s like taking apart a common household appliance to see how everything fits together. Now we can understand how the proteins in the complex come together to perform their function,” says Ryan Abdella, the paper’s corresponding first author.
According to Roger Kornberg, a prominent biochemist and professor of structural biology at Stanford University School of Medicine, it has been near impossible to view the tiny molecular structure. “These complexes are scarce. It takes hundreds of liters of human cells, which are very hard to grow, to obtain small amounts of the protein complexes,” he explains.
He and colleagues placed a tissue sample atop a single sheet of graphene oxide which reduced the quantity of tissue required for scanning and enhanced the image produced. Researchers then applied cryo-EM to identify the three-dimensional form of Mediator. This method uses a beam of electrons to capture several 2D pictures of microscopic substances.
“Some of those subunits were already known from other experiments, but we had no idea how the pieces assembled together or interacted with each other. With our final structure, we were finally able to see this whole complex and understand its organization,” says co- corresponding first author.Anna Talyzina, a graduate student of He’s lab.
The image revealed the Mediator as a flattened protein complex that is approximately 45 nanometers long. It is able to attach to the transcription factors (namely RNA Polymerase II) toward the end of the molecule, a discovery that astonished the researchers. As a result, “Mediator moves like a pendulum,” says Abdella. “Next, we want to understand what this flexibility means. We think it might have an impact on the activity of a key enzyme within the complex.”
This study is published in NCBI.